Polyacrylamide Gel Electrophoresis
Overview
Polyacrylamide gel electrophoresis (PAGE) is a method of separating DNA fragments by length. DNA fragments are loaded into a gel made of many acrylamide polymers. A voltage differential is applied across the gel, causing the negatively charged DNA fragments to move down the gel. Shorter fragments will encounter less resistance from the gel matrix and travel further than longer fragments in the same amount of time. The gel is later stained with a fluorescent DNA binding dye so that the DNA can be seen as bands in a blue light or UV imager. Typically, a DNA ladder with DNA fragments of known size is loaded into the gel as a "ruler" for samples to be compared against.
The higher the concentration of the gel, the slower all DNA travels through it, enabling a higher resolution to be obtained. Single stranded DNA will travel faster than double stranded DNA. Plasmid (circularized) DNA will travel faster on a gel and appear shorter than linear DNA of the same length.
Single Nucleotide Addition Detection
By adding desired nucleotides one base at a time using Enzymatic Synthesis Methods, a completely arbitrary sequence of DNA can be created without an existing template sequence. SSI Biology uses PAGE to detect addition of exactly one nucleotide to an existing strand of single-stranded DNA. We currently use 15% pre-cast gels and SYBR Gold gel stain for imaging the DNA.
Safety
Polyacrylamide is a neurotoxin when in liquid form. However, this is only a concern when casting gels from liquid polyacrylamide. We only use pre-cast polyacrylamide gels, which are delivered in solid form and are non-hazardous.
Procedure
Setting up the gel apparatus

SSI Biology currently uses the Bio-Rad Mini PROTEAN Tetra Cell apparatus.
Take a 15% TBE-Urea denaturing pre-cast gel out of its package, remove the green tape from the bottom of the gel, and carefully remove the comb from the top of the gel. Slot the gel and a buffer dam into the gel apparatus and push the green wings inwards, making sure there are no gaps where the gel meets the bottom tabs of the apparatus and that the right sides of the gel and the buffer dam are facing outwards. A second gel can be placed into the gel apparatus instead of the buffer dam. Place the gel apparatus into its clear box, making sure that the positive (red) electrode is closer to the red mark on the box. Fill the apparatus with .5x Tris-Borate-EDTA (TBE) running buffer, stopping to seal any leaks. Let the buffer overflow into the gel box up to the level marked on the box. Up to four gels can be run at once in the same box. Pipette TBE buffer up and down into the each well of the gel to rinse out any urea residue. Place the green lid onto the box and pre-run the gel for 20 minutes at your desired voltage.
Loading samples
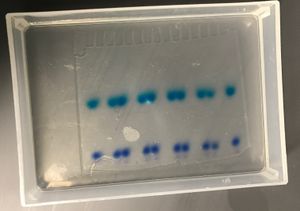
Prepare your DNA samples by mixing them with 2x Gel Loading Buffer II (containing the denaturing agent formamide and bromophenol blue and xylene cyanol dyes). Take the lid off the gel box, and rinse the wells with TBE buffer before loading your samples. Pipette each sample carefully into one well, so as to avoid bubbles or overflow into other wells. Cover the gel box with the green lid and connect the lid to the power supply. Run the gel at 150V until the dark blue dye (the first dye front) gets to about 1 cm from the bottom of the gel.
Staining the gel
Turn off the power supply, open the gel box, and disassemble the gel apparatus. Carefully pry the gel apart with a lever or spatula. Slide the gel from the plastic cover into a tray with 100 uL 1x SYBR Gold gel stain. Let the gel sit in the staining solution for 30 minutes.
Imaging the gel
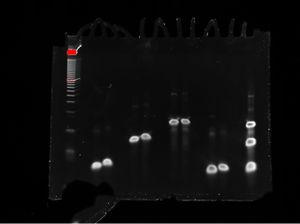
Carry the gel tray to the gel imager and carefully place the gel into the imager, centering it on the screen. Set the imager to optimize for faint bands and SYBR Gold staining, and be sure to save the image of the gel that appears.
Example Procedure
An example of a PAGE procedure is linked here.
