Pyrosequencing
Pyrosequencing is a sequencing technique that uses a four-enzyme pathway to detect the addition of nucleotides to a strand of DNA. Pyrosequencing is compatible with Electrowetting on Dielectric microfluidics systems.
Overview
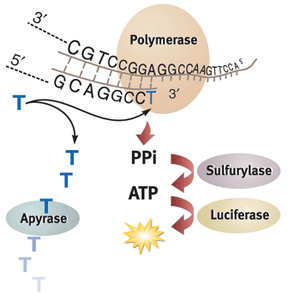
In commercial pyrosequencing, the strand to be sequenced is immobilized on magnetic beads and used as a template for a template-dependent polymerase. One type of nucleotide at a time is introduced to the DNA with enzyme mix to build up the complementary sequence to this strand.
1. When the correct type of nucleotide is added onto the complementary DNA strand by a template-dependent polymerase, pyrophosphate (PPi) is released as a byproduct of the DNA backbone formation reaction.
2. The reaction of this pyrophosphate and adenosine phosphosulfate (APS) to form ATP is then catalyzed by ATP sulfurylase.
3. The ATP produced then powers the conversion of luciferin substrate into a luminescence signal almost instantaneously by firefly luciferase. The light signal resulting from each nucleotide addition lasts about 10 minutes and can be measured by a photomultiplier tube.
4. Pyrosequencing can be sped up with the addition of apyrase, which degrades luminescence signal quickly and allows for the next nucleotide to be added. Alternatively, unreacted reagents and nucleotides can be washed off the DNA before the introduction of the next nucleotide type.
If luminescence is detected, the type of nucleotide that was added is recorded, and the process of adding nucleotides is continued until the full sequence of the complementary strand is known.
To control the reaction steps, pyrosequencing reagents are typically separated into enzyme mix (containing ATP sulfurylase, polymerase, luciferin, and luciferase) and substrate mix (containing APS and one type of nucleotide). The nucleotide 2'-Deoxyadenosine-5'-O-(1-Thiotriphosphate), or dATPαS, is used in place of dATP in pyrosequencing reactions since dATP is a substrate for luciferase and will give a false positive luminescence signal. Luminescence signal is linearly proportional to the number of nucleotides added (up to about 4 or 5 bases), so a sequence of two or more of the same type of nucleotide in a row (a homopolymer) can be distinguished from addition of a single base.
The Qiagen Pyromark system and the [Roche 454 Sequencing]
Modified Pyrosequencing
SSI Biology uses a modified version of pyrosequencing to quickly detect extension of a starting single strand of DNA instead of the sequence of a strand. TdT or another of our Enzymatic Synthesis Methods is used to add a known base instead of polymerase. The pyrophosphate from this addition can then be converted to a luminescence signal and detected. Luminescence indicates successful addition of one or more bases to the DNA. By comparing signal level to signal produced by a control addition reaction with blocked bases, we can also determine whether an insertion error occurred (more than one base was added).
Sample Procedure
A starting oligonucleotide of DNA is extended with a desired nucleotide and one of the Enzymatic Synthesis Methods listed above. The solution containing pyrophosphates (this can be the unpurified extended DNA sample or the waste after the DNA is purified) is combined with 3x enzyme mix (13.5 mU/uL ATP Sulfurylase, 1.5 ug/uL luciferase, and 1.8 ug/uL luciferin in pyrosequencing buffer) and 3.09x substrate mix (7.5 ug/uL APS in pyrosequencing buffer) in the correct dilution for a total sample volume of 20 uL. A reducing agent such as dithiothreitol or TCEP can be added but is not necessary for the reaction to occur. The samples should be mixed thoroughly and measured as soon after adding enzyme mix as possible. Samples are transferred to a 96-well plate for measurement and loaded into a plate reader, which will read out luminescence values for the sample over a given time period.
An example protocol for a basic pyrosequencing experiment can be found here.
References
Boles, Deborah J., et al. "Droplet-based pyrosequencing using digital microfluidics." Analytical chemistry 83.22 (2011): 8439-8447. [1]
